|
12/12/2023 0 Comments Klenow fragment pdf 
coli genomic DNA using oligonucleotide primers corresponding to the 16S rRNA locus. coli is cleaved by the protease enzyme subtilisin. coli contamination is evaluated using 5 µL replicate samples of enzyme solution that are denatured and screened in a TaqMan qPCR assay for the presence of contaminating E. The Klenow fragment is a large protein fragment which is produced when DNA polymerase I from E. Single-stranded exonuclease is determined in a 50 µL reaction containing a radiolabeled single-stranded DNA substrate and 10 µL of enzyme solution incubated for 4 hours at 37☌.ĭouble-stranded exonuclease activity is determined in a 50 µL reaction containing a radiolabeled double-stranded DNA substrate and 10 µL of enzyme solution incubated for 4 hours at 37☌.ĭouble-stranded endonuclease activity is determined in a 50 µL reaction containing 0.5 µg of plasmid DNA and 10 µL of enzyme solution incubated for 4 hours at 37☌.Į. The fragment still has the polymerase and 35 exonuclease activity, but lacks the 53 exonuclease activity of the holoenzyme (). Purity is assessed by comparing the aggregate mass of contaminant bands in the concentrated sample to the band's mass corresponding to the protein of interest in the diluted sample. The Klenow fragment of DNA polymerase I is a proteolytic fragment obtained by the treatment of DNA polymerase I with subtilisin. Physical purity is evaluated by SDS-PAGE of concentrated and diluted enzyme solutions followed by silver-stain detection. Use of Fluorescence Resonance Energy Transfer To Investigate the Conformation of DNA Substrates Bound to the Klenow Fragment. Protein concentration is determined by OD 280 absorbance. The fluorescent oligomer bound tightly to the Klenow fragment (KD 7.9 nM), and the probes position within. They have used a technique called RNAprimed, array-based Klenow enzyme assay because it involves Klenow fragment of DNA polymerase I. Reactions were incubated for 10 minutes at 37☌, placed on ice and analyzed using the method of Sambrook and Russel (Molecular Cloning, v3, 2001, pp. 
Dilutions of the enzyme were made in a 50% glycerol Klenow (3ʹ→5ʹ exo–) storage solution and added to 50 µL reactions containing calf thymus DNA, 1x Klenow Reaction Buffer, 3H-dTTP and 100 µM dNTPs. 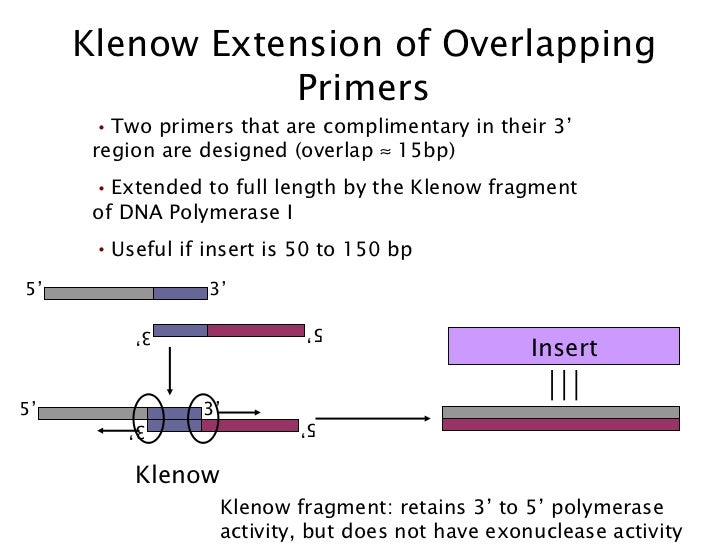
Unit activity was measured using a two-fold serial dilution method. Stop the reaction by adding EDTA (final concentration of 10mM) and heating at 75☌ for 20 minutes. Incubate the reaction mixture at 25☌ for 15 minutes.ģ. Note: Klenow Fragment, Exonuclease Minus, will leave a single-base 3´-overhang for a significant proportion of the DNA fragments during the fill-in reaction (8). Stop the reaction by heating the mixture for 10 minutes at 75☌. Incubate the reaction at room temperature for 10 minutes. Add 1 U of Klenow Fragment per microgram of DNAĢ. unit of Klenow Fragment per microgram of DNA.Add dNTPs (each at a final concentration of 33 µM). 
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
